
Barnum Family DNA Project
![]()
–Remember, DNA is the Gene in Genealogy –
Project Surnames:
|
BARNUM BARNHAM BARNAM |
Introduction to the Barnum Family DNA Project
You’ve seen on the welcome page that it is now possible to confirm a link where no conventional source records exist. You may wonder, though, how we can do that.
First, a painless cheek scraping is used to obtain a DNA sample that, through
Y-DNA genetic sequencing, can accurately either confirm a relationship with a
99.9-percent probability or provide a 100-percent certainty that no near-term
relationship existed. The primary test attempts to determine if two
individuals, whether thought to be related or unrelated, actually did have a
common ancestor. A match is best done by testing a male member of a known
biological family (brother, father or first cousin) against the potential male
relative.
[The female side of the family can also be tested,
through mitochondrial DNA (MtDNA) from daughter to mother to grandmother, etc.
Click here
for a few Y-DNA and MtDNA success stories.]
Y chromosome DNA, passed from father to son, and mitochondrial DNA, passed from
mother to daughter, are varied through the generations only by occasional
natural mutations called markers. These mutations, occurring in an otherwise
continuous string of genetic replication, serve as genetic signposts for
tracing human evolution. By following a marker back through time to its origin,
geneticists can identify the Most Recent Common Ancestor (MRCA) of everyone
alive who carries a common marker. The divergent branches of the human family
tree, represented by groups carrying a given marker, can be followed back to
“nodes” on the tree where a mutation split a branch into two directions.
With appropriate analysis, we can now get a pretty good idea of who the shared genetic ancestor was. One of our goals is to find the time to the most recent common ancestor (TMRCA) of the various branches of the Barnum family. Scientists can calculated that time based upon the observed number of mutations by which two Y chromosomes differ. That is because mutations occurring at random can be fit into a probability distribution function.
The approach for the Barnum Family DNA Project will be to develop groups of matching DNA sequences and to cross reference that information against the oldest known ancestor. For now, that person is the immigrant ancestor, Thomas Barnum. My hope is that we can find more conclusive evidence about the supposed English origins of Thomas and the countries of origin of his more-distant ancestors, as well as helping to connect “orphan” Barnum families to the larger family tree.
Step 1
The first thing we need to do is to determine which incoming samples belong to each family group (for example, descendants of each of the four sons of the immigrant ancestor) and which remain un-matched. The easiest and quickest approach is to simply add the sums of the differences between the Y-DNA values for each tested family member. The larger the value for the sum of the differences, the farther each group is separated.
Step 2
As family groups are found, they will be arranged together on the “results” page. (The results page will be updated as new results arrive with a relationship to existing groups.)
Step 3
When enough samples have been received, other equations can be applied. Usually, the serious work can be performed when we have at least 40 samples. That is when we can start reliable calculations for the Most Recent Common Ancestor (MRCA). As stated earlier, for now that will be the immigrant ancestor Thomas Barnum. Over time, however, we should begin to see data pointing to our first Paleo ancestor to enter Europe.
Step 4
By this time, we should have a pretty good idea of how many Barnum immigrant lines have survived by repeating the iterations of groupings. We also should begin to see two new items: Time to Most Recent Common Ancestor (TMRCA) and general country location of the Most Recent Common Ancestor (by the use of haplogroup studies).
Within Step 4, I should be able to begin posting Phylogenetic graphs of the Barnum Family DNA Project data. (Phylogenetic graphs are those based on common evolutionary descent and recency of common ancestry).
For the more curious, here are a few terms that you may see used in genetic genealogy. If some of these terms seem esoteric or overly scientific, don’t despair. As you become more involved in genetic genealogy they’ll become familiar and more readily understandable:
The Y Chromosome contains the unique feature that
causes a person to be male and it is only transmitted from fathers to their
sons. The Y Chromosome is virtually identical through a number of generations
over time. However, rare mutations can occur.
Time to Most Recent Common Ancestor is calculated by using DNA sequences
showing variation in the population (also called genetic markers or simply
markers) to provide information on how closely the Y chromosomes from two
individuals are related. Relatedness is quantified by TMRCA, the Time to Most
Recent Common Ancestor, which is stated as the number of generations between
two examined Y chromosomes and a common ancestor. Estimates of TMRCA are based
on the observed number of mutations by which the two Y chromosomes differ.
Since mutations occur at random, the estimate of a TMRCA is not an exact number
(i.e., 7 generations), but rather a probability distribution. As one uses more
and more markers, the distribution becomes tighter and tighter about its mean
value (its variance becomes smaller), and estimates have higher precision.
SNPs (pronounced “Snips”) stands for Single Nucleotide Polymorphisms. The problem with SNPs for genealogical work is that they have very low mutation rates. This makes them useful for very deep genealogies (i.e. thousands of generations), but not too helpful for genealogists because they are not recent enough.
Below is an example of a haplogroup tree, showing a number of Y chromosome markers (SNPs and subclades) that may be used to differentiate different lines of descent:
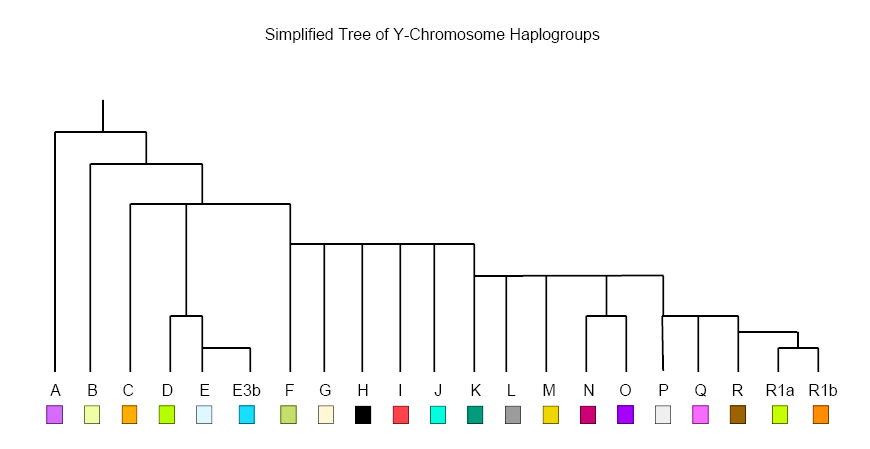
A type of DNA sequence called a microsatellite has a very high mutation
rate, so that it will only stay in its current state for a few generations
(with mutations rates on the order of 1:500). Microsatellites are small blocks
of DNA with repeated sequences, for example ATATATAT is a sequence with four AT
repeats.
DYS# (DNA Y-Chromosome Segment marker number) is a label for
genetic markers on the Y chromosome. Each marker is designated by a number,
according to international conventions. At present, most of the DYS
designations in use are given to STR markers (Short Tandem Repeats – a
microsatellite class often used in genetic genealogy).
Virtually all times reported in genetic genealogy are generation times. One can
translate these into years back to a common relative by making assumptions
about the average number of years in a standard human generation. The problem
here is that the values for a generation in the literature are not standard,
ranging from 15 to 25 years.
Mutation Rate Boundaries for the TMRCA Calculator used by Family Tree DNA (FTDNA):
1) Standard – assumes that the
mutation rate per marker (u) is u =1:500 = 0.002. This is the average of a
number of studies. In other words, we can expect any one marker to mutate once
in 500 generations.
2) High-u = 0.004. This places a lower boundary on
TMRCA, that is, these times are likely underestimates of the true times.
What is the effect of adding even more markers (increasing from 12 to 25 or from 25 to 37, for example? The answer is that adding additional markers increases the precision of the test.
| DNA PROJECT | HOME | EMAIL | SURNAMES |
![]()
A Research Guide to the Genealogy of the Barnum/Barnam/Barnham Family Worldwide
![]()
The information on this site is developed and maintained by Patrick Barnum

©1998, 2022. The format of this website and all original statements and narrative included on it are copyrighted and all rights are reserved. Factual information may be freely quoted for use in private genealogical research when accompanied by a full source citation, including the date of acquisition. Click here to view the format of a citation for an Internet resource. The publication of large extracts from this site in any form requires prior written consent.